| Download the amazing global Makindo app: ✅ Means NICE/National Guidelines 2026 compliant Android | Apple | |
|---|---|
| MEDICAL DISCLAIMER: Educational use only. Not for diagnosis or management. See below for full disclaimer. |
DNA and RNA
Related Subjects: |DNA and RNA short notes |DNA replication |DNA structure in Nucleus |Mitosis and Meiosis |Telomeres |Autosomal Recessive |X Linked Recessive |Autosomal Dominant |Li Fraumeni syndrome |Genetic Linkage |Cell Cycle
Chargaff's rules: In double-stranded DNA ➡️ %A = %T and %G = %C. Purines = Pyrimidines overall. These base-pairing rules were known before Watson & Crick’s helix model.
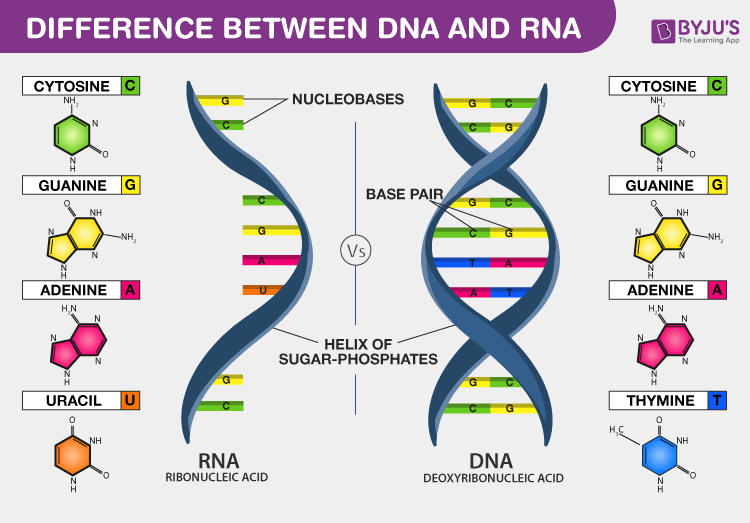
🧱 Building Blocks
- Purines (2 rings): Adenine (A), Guanine (G). Synthesis requires glycine, aspartate, glutamine.
- Pyrimidines (1 ring): Cytosine (C), Thymine (T in DNA), Uracil (U in RNA).
- Base pairing → Purine + Pyrimidine (3 rings across) maintains uniform helix width.
📜 Nucleic Acids
- DNA: A=T (2 H-bonds), G≡C (3 H-bonds - stronger!).
- RNA: A=U, G≡C. Usually single-stranded.
- Sugars: DNA = deoxyribose; RNA = ribose (extra OH → less stable).
- Human diploid genome = 23 chromosome pairs.
- DNA → nuclear; RNA → 90% cytoplasmic.
🔬 Types of RNA
- mRNA: Protein template. 5′ cap + 3′ poly(A) tail = stability & translation.
- tRNA: Cloverleaf adaptor carrying amino acids (anticodon matches codon).
- rRNA: Structural + catalytic core of ribosomes (peptidyl transferase activity).
📆 Eukaryotic Cell Cycle
- G0: Resting phase (neurons, RBCs stay here permanently).
- G1: Growth, organelle duplication. Restriction point checks DNA damage.
- S: DNA replication. Centrosome duplication.
- G2: Error checking before mitosis.
- M: Mitosis (prophase → telophase + cytokinesis).
⚡ DNA Replication
- Semiconservative: 1 parental + 1 new strand.
- DNA polymerase: works 5′→3′; needs an RNA primer.
- Leading strand: continuous synthesis.
- Lagging strand: Okazaki fragments (later ligated).
- Error rate: ~1 per 10¹⁰ bases (proofreading via 3′→5′ exonuclease).
- mtDNA: circular, maternal inheritance.
- Proto-oncogenes regulate cycle; mutations → oncogenes → cancer.
🔄 Quality Control
- Checkpoints (cyclins + CDKs regulate):
- G1/S: “restriction point” – damaged DNA? nutrients sufficient?
- G2/M: DNA replicated? errors fixed?
- Metaphase checkpoint: spindle attachment?
⏳ Telomeres
- Repeat sequences (TTAGGG in humans) protect chromosome ends.
- Shorten with each division → cellular aging (“replicative senescence”).
- Germline, stem, and cancer cells express telomerase → immortality.
⚡ Energy for Replication
- Energy comes from hydrolysis of nucleoside triphosphates → pyrophosphate release drives chain elongation.
- Strand growth always at the 3′ end.
- Diagram: Okazaki fragments
📌 Quick Exam Summary
🧬 A=T (2 bonds), G≡C (3 bonds). 📆 Cell cycle checkpoints stop faulty replication. ⚡ Replication is semiconservative, 5′→3′, with leading vs lagging strands. ⏳ Telomeres shorten with age unless telomerase is active. 💥 Oncogenes = mutated proto-oncogenes → unchecked growth.
Categories
- A Level
- About
- Acute Medicine
- Anaesthetics and Critical Care
- Anatomy
- Anatomy and Physiology
- Biochemistry
- Book
- Cardiology
- Collections
- CompSci
- Crib Sheets
- Dental
- Dermatology
- Differentials
- Drugs
- ENT
- Education
- Electrocardiogram
- Embryology
- Emergency Medicine
- Endocrinology
- Ethics
- Foundation Doctors
- GCSE
- Gastroenterology
- General Practice
- Genetics
- Geriatric Medicine
- Guidelines
- Gynaecology
- Haematology
- Hepatology
- Immunology
- Infectious Diseases
- Infographic
- Investigations
- Lists
- MRCP
- Mandatory Training
- Medical Students
- Microbiology
- Nephrology
- Neu
- Neurology
- Neurosurgery
- Nutrition
- OSCE
- OSCEs
- Obstetrics
- Obstetrics Gynaecology
- Oncology
- Ophthalmology
- Oral Medicine and Dentistry
- Orthopaedics
- Paediatrics
- Palliative
- Pathology
- Pharmacology
- Physiology
- Procedures
- Psychiatry
- Public Health
- Radiology
- Renal
- Respiratory
- Resuscitation
- Revision
- Rheumatology
- Statistics and Research
- Stroke
- Surgery
- Toxicology
- Trauma and Orthopaedics
- USMLE
- Urology
- Vascular Surgery
